absDL
In this repository we arranged the required elements for training a deep network to deploy single-shot absorption imaging in your lab.
We provide a Matlab script to assist with correctly handling the training and prediction images, and a python script for the network training. Both will enable you to reproduce the illustration figures from the paper:
Single-exposure absorption imaging of ultracold atoms using deep learning (Phys. Rev. Applied 14, 014011 (2020)) (arXiv:2003.01643)
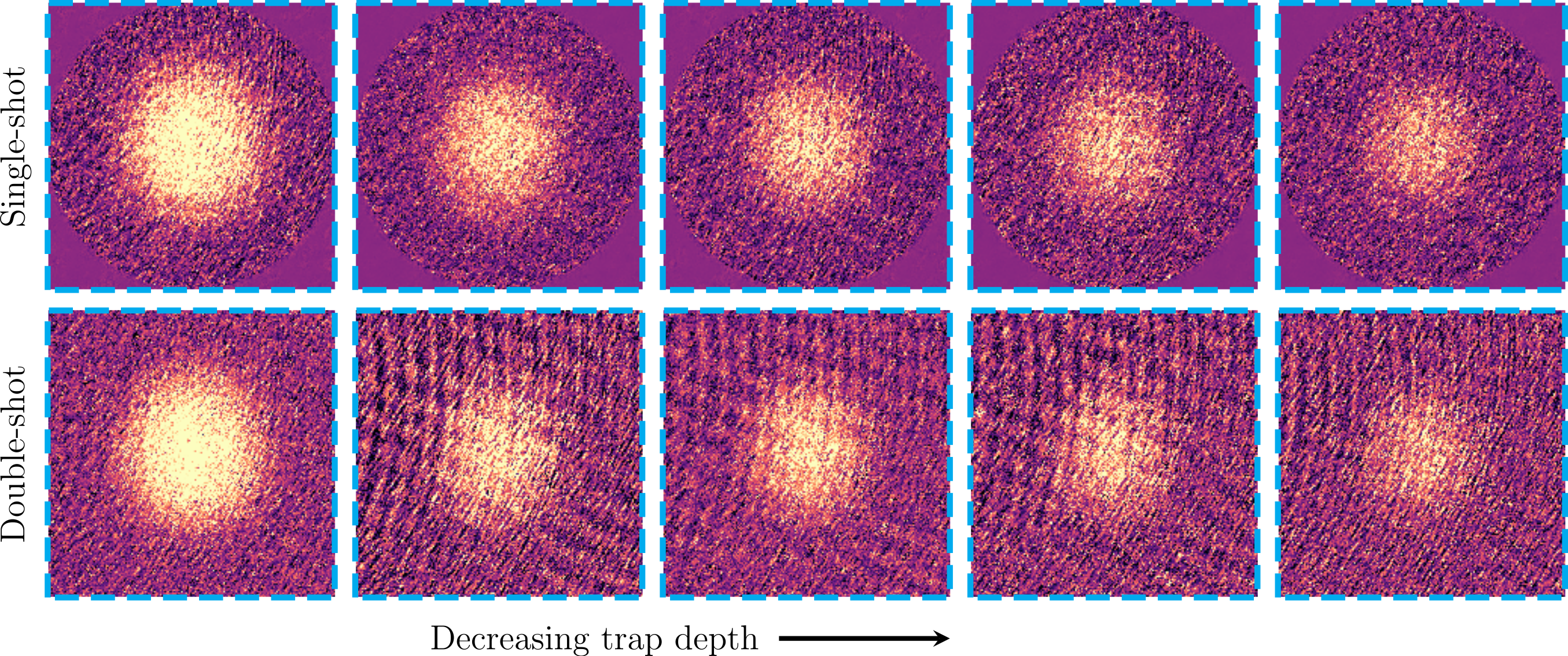
Here we provide hyperparameters and architecture that were used to generate the results in the paper. Most of them were crudely optimized for the Technion ultracold fermions lab apparatus and will require some tunning to fit other use-cases. We encourage you to share your insights!
absDL repository is provided and maintained by Yoav Sagi’s ultracold fermions research group from the Technion – Israel Institute of Technology. You are very welcome to use and contribute to it!
Environment:
1. Using Anaconda recipe:
Please follow the Anaconda recipe provided with this repo. Download from here.
2. Using pip:
- make sure that your virtual environment is installed with python 3.7 or 3.6.8
- make sure your pip is pointing on your venv interpreter: <venv_location/bin/pip>
- on the project directory, type in:
$ pip install -r requirements.txt
Dataset:
Use the Matlab script ‘dataset_extractor.m’ to extract a 32bit TIFF images from your experimental setup. In case you have the raw camera images stored as MAT files small modifications to this script would do. Even if it is not the case, you can follow its lines to understand how we normalized the images and extract training and testing sets for your data.
Original dataset:
The dataset used in the paper can be downloaded from zenodo. We use constant normalization for all frames to adjust the log images and store them in separate TIFF files. In our experimental setup the camera is installed from beneath the vacuum chamber. Therefore, to focus the image on the free-falling atoms, the imaging system was manually shifted to follow the gravitational- induced position of the atoms. This introduced a variation of the noise patterns between sets of images, which was otherwise quite limited. A small (10MBs) sample from our dataset is provided directly in the GitHub repository, under /miniset/.
Training:
You can simply run ‘main.py’ after referencing the datasets with and without atoms in ‘wAtoms_ds.txt’ and ‘woAtoms_ds.txt’, respectively. The main script includes an argument parser to allow variation of the options and parameters from within the terminal. You can view the different options by running main.py with the -h flag (or –help).